Descriptive vs. inferential community detection
Community detection is the task of dividing a network — typically one which is large — into many smaller groups of nodes that have a similar contribution to the overall network structure. With such a division, we can better summarize the large-scale structure of a network by describing how these groups are connected, instead of each individual node. This simplified description can be used to digest an otherwise intractable representation of a large system, providing insight into its most important patterns, how they relate to its function, and the underlying mechanisms responsible for its formation.
At a very fundamental level, community detection methods can be divided into two main categories: “descriptive” and “inferential.”
Descriptive methods attempt to find communities according to some context-dependent notion of a good division of the network into groups. These notions are based on the patterns that can be identified in the network via an exhaustive algorithm, but without taking into consideration the possible rules that were used to create them. These patterns are used only to describe the network, not to explain it. Usually, these approaches do not articulate precisely what constitutes community structure to begin with, and focus instead only on how to detect them. For this kind of method, concepts of statistical significance, parsimony and generalizability are usually not evoked.
Inferential methods, on the other hand, start with an explicit definition of what constitutes community structure, via a generative model for the network. This model describes how a latent (i.e. not observed) partition of the nodes would affect the placement of the edges. The inference consists on reversing this procedure to determine which node partitions are more likely to have been responsible for the observed network. The result of this is a “fit” of a model to data, that can be used as a tentative explanation of how it came to be. The concepts of statistical significance, parsimony and generalizability arise naturally and can be quantitatively assessed in this context. See e.g. [2].
Descriptive community detection methods are by far the most numerous, and those that are in most widespread use. However, this contrasts with the current state-of-the-art, which is composed in large part of inferential approaches. Here we point out the major differences between them and discuss how to decide which is more appropriate, and also why one should in general favor the inferential varieties whenever the objective is derive interpretations from data.
Describing vs. explaining
We begin by observing that descriptive clustering approaches are the method of choice in certain contexts. For instance, such approaches arise naturally when the objective is to divide a network into two or more parts as a means to solve a variety of optimization problems. Arguably, the most classic example of this is the design of Very Large Scale Integrated Circuits (VLSI). The task is to combine millions of transistors into a single physical microprocessor chip. Transistors that connect to each other must be placed together to take less space, consume less power, reduce latency, and reduce the risk of cross-talk with other nearby connections. To achieve this, the initial stage of a VLSI process involves the partitioning of the circuit into many smaller modules with few connections between them, in a manner that enables their efficient spatial placement, i.e. by positioning the transistors in each module close together and those in different modules farther apart.
Another notable example is parallel task scheduling, a problem that appears in computer science and operations research. The objective is to distribute processes (i.e. programs, or tasks in general) between different processors, so they can run at the same time. Since processes depend on the partial results of other processes, this forms a dependency network, which then needs to be divided such that the number of dependencies across processors is minimized. The optimal division is the one where all tasks are able to finish in the shortest time possible.
Both examples above, and others, have motivated a large literature on “graph partitioning” dating back to the 70s, which covers a family of problems that play an important role in computer science and algorithmic complexity theory.
Although reminiscent of graph partitioning, and sharing with it many algorithmic similarities, community detection is used more broadly with a different goal [3]. Namely, the objective is to perform data analysis, where one wants to extract scientific understanding from empirical observations. The communities identified are usually directly used for representation and/or interpretation of the data, rather than as a mere device to solve a particular optimization problem. In this context, a merely descriptive approach will fail at giving us a meaningful insight into the data, and can be misleading, as we will discuss in the following.
We illustrate the difference between descriptive and inferential approaches in Figure 1. We first make an analogy with the famous “face” seen on images of the Cydonia Mensae region of the planet Mars. A merely descriptive account of the image can be made by identifying the facial features seen, which most people immediately recognize. However, an inferential description of the same image would seek instead to explain what is being seen. The process of explanation must invariably involve at its core an application of the law of parsimony, or Occam's razor. This principle predicates that when considering two hypotheses compatible with an observation, the simplest one must prevail. Employing this logic results in the conclusion that what we are seeing is in fact a regular mountain, without denying that it looks like a face in that picture, but just accidentally. In other words, the “facial” description is not useful as an explanation, as it emerges out of random features rather than exposing any underlying mechanism.
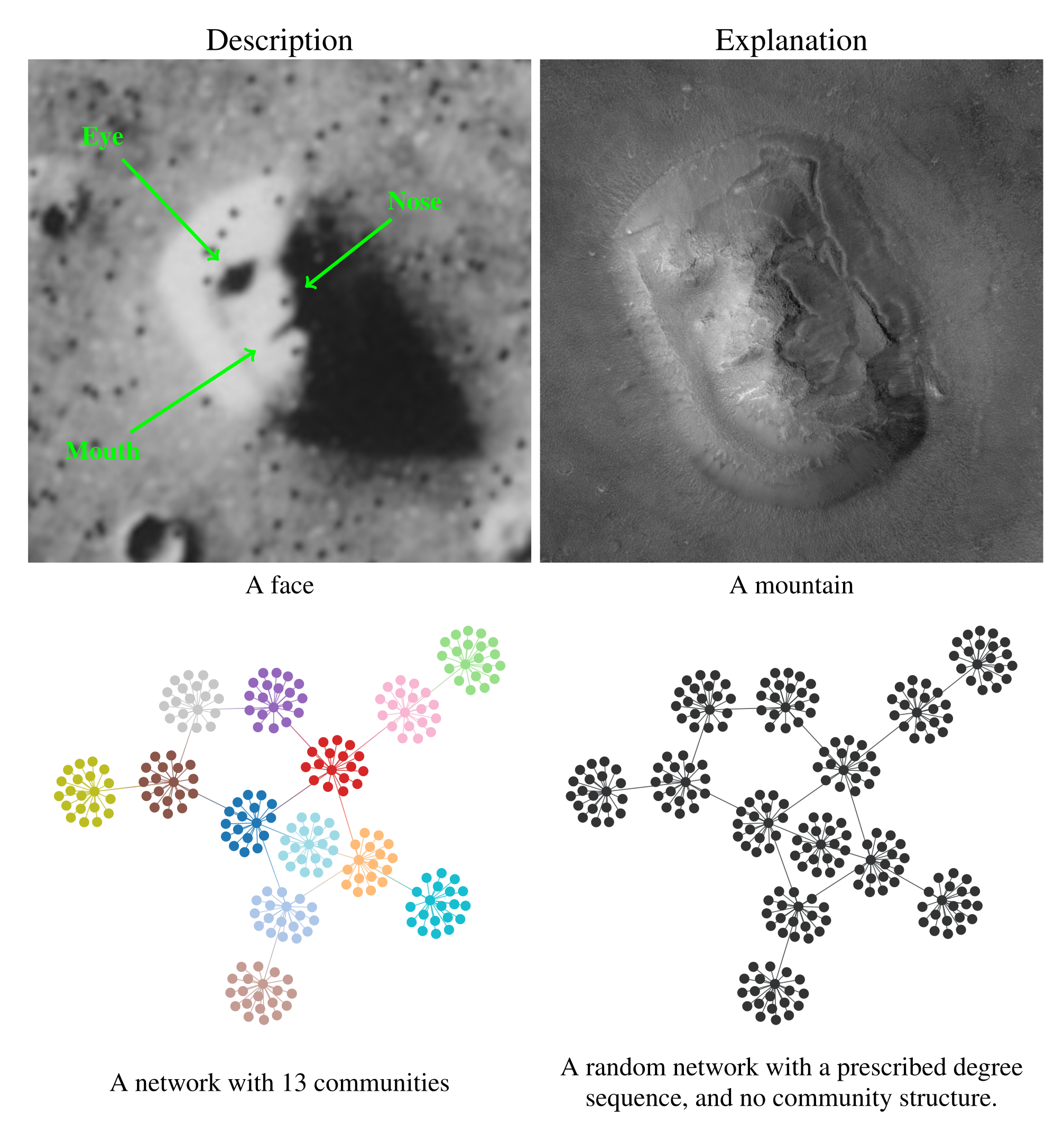
Going out of the analogy and back to the problem of community detection, in the bottom of Figure 1 we see a descriptive and an inferential account of an example network. The descriptive one is a division of the nodes into 13 assortative communities, which would be identified with many descriptive community detection methods available in the literature. Indeed, we can inspect visually that these groups form assortative communities, and most people would agree that these communities are really there, according to most definitions in use: these are groups of nodes with many more internal edges than external ones. However, an inferential account of the same network would reveal something else altogether. Specifically, it would explain this network as the outcome of a process where the edges are placed at random, without the existence of any communities. The communities that we see in Figure 1 (a) are just a byproduct of this random process, and therefore carry no explanatory power. In fact, this is exactly how the network in this example was generated, i.e. by choosing a specific degree sequence and connecting the edges uniformly at random.
In Figure 2 (a) we illustrate in more detail how the network in Figure 1 was generated: The degrees of the nodes are fixed, forming “stubs” or “half-edges”, which are then paired uniformly at random forming the edges of the network. In Figure 2 (b), like in Figure 1, the node colors show the partition found with descriptive community detection methods. However, this network division carries no explanatory power beyond what is contained in the degree sequence of the network, since it is generated otherwise uniformly at random. This becomes evident in Figure 2 (c), where we show another network sampled from the same generative process, i.e. another random pairing, but partitioned according to the same division as in Figure 2 (b). Since the nodes are paired uniformly at random, constrained only by their degree, this will create new apparent “communities” that are always uncorrelated with one another. Like the “face” on Mars, they can be seen and described, but they cannot explain.
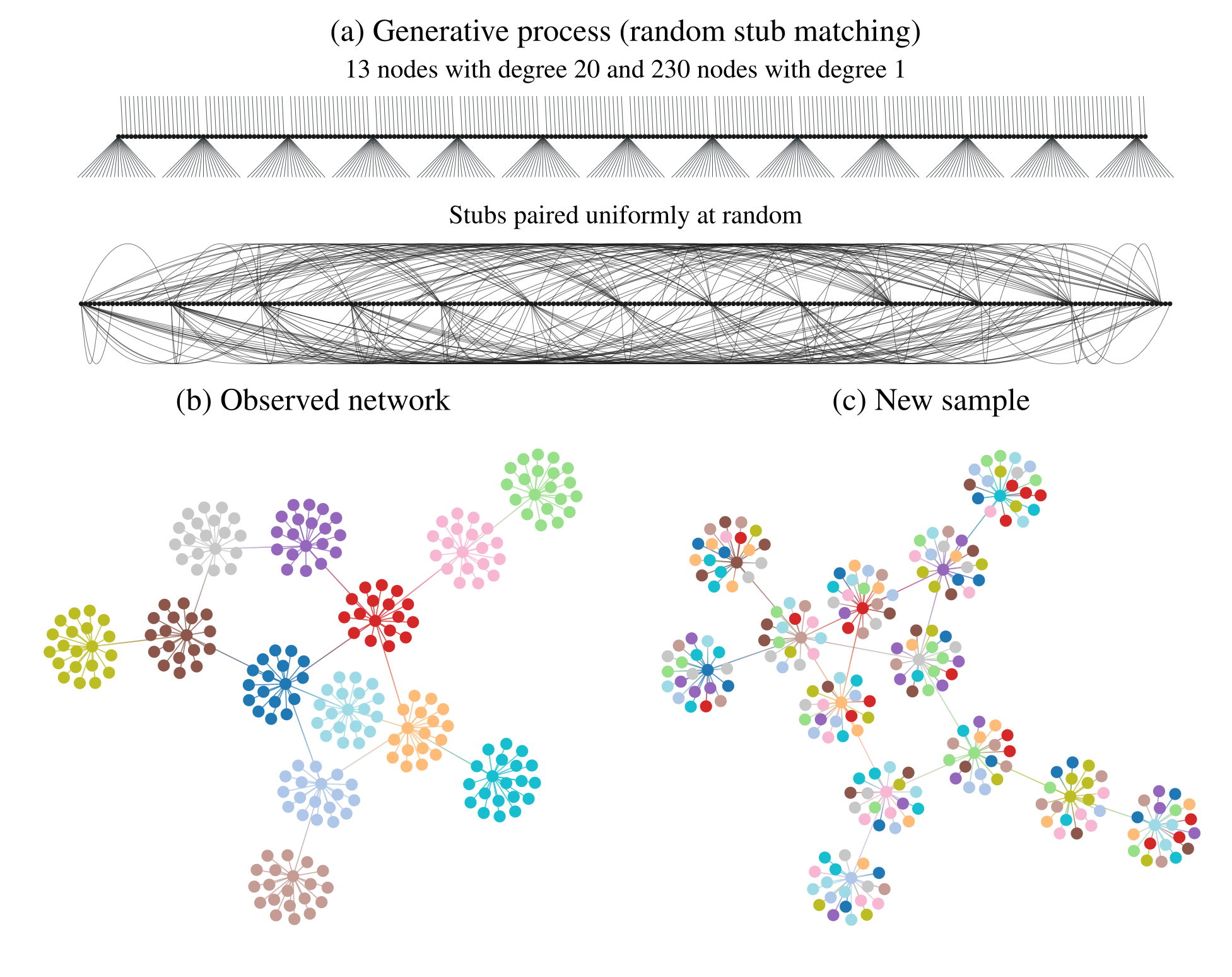
We emphasize that the communities found in Figure 2 (b) are indeed really there from a descriptive point of view, and they can in fact be useful for a variety of tasks. For example, the cut given by the partition, i.e. the number of edges that go between different groups, is only 13, which means that we need only to remove this number of edges to break the network into (in this case) 13 smaller components. Depending on context, this kind of information can be used to prevent a widespread epidemic, hinder undesired communication, or, as we have already discussed, distribute tasks among processors and design a microchip. However, what these communities cannot be used for is to explain the data. In particular, a conclusion that would be completely incorrect is that the nodes that belong to the same group would have a larger probability of being connected between themselves. As shown in Figure 2 (a), this is clearly not the case, as the observed “communities” arise by pure chance, without any preference between the nodes.
To infer or to describe? A litmus test
Given the above differences, and the fact that both inferential and descriptive approaches have their uses depending on context, we are left with the question: Which approach is more appropriate for a given task at hand? In order to help answering this question, independent of the particular context, it is useful to consider the following “litmus test”:
Q: “Would the usefulness of our conclusions change if we learn, after obtaining the communities, that the network being analyzed is completely random?”
If the answer is “yes”, then an inferential approach is needed.
If the answer is “no”, then an inferential approach is not required.
If the answer to the above question is “yes”, then an inferential approach is warranted, since the conclusions depend on an interpretation of how the data were generated. Otherwise, a purely descriptive approach may be appropriate since considerations about generative processes are not relevant.
It is important to understand that the relevant question in this context is not whether the network being analyzed is actually fully random, 1 since this is rarely the case for empirical networks. Instead, considering this hypothetical scenario serves as a test to evaluate if our task requires us to separate between actual latent community structure (i.e. those that are responsible for the network formation), from those that arise completely out of random fluctuations, and hence carry no explanatory power. Furthermore, most empirical networks, even if not fully random, like most interesting data, are better explained by a mixture of structure and randomness, and a method that cannot tell those apart cannot be used for inferential purposes.
1 “Fully random” here means sampled form a random graph model, like the Erdős-Rényi model, the configuration model, or some other null model where whatever communities we may ascribe to the nodes play no role in the placement of the edges.
2 Although this is certainly true at a first instance, we can also argue that properly understanding why a certain partition was possible in the first place would be useful for reproducibility and to aid the design of future instances of the problem. For these purposes, an inferential approach would be more appropriate.
Returning to the VLSI and task scheduling examples we considered in the previous section, it is clear that the answer to the litmus test above would be “no”, since it hardly matters how the network was generated and how we should interpret the partition found, as long as the integrated circuit can be manufactured and function efficiently, or the tasks finish in the minimal time. Interpretation and explanations are simply not the primary goals in these cases.2
However, it is safe to say that in network data analyses very often the answer to the question above question would be “yes.” Typically, community detection methods are used to try to understand the overall large-scale network structure, determine the prevalent mixing patterns, make simplifications and generalizations, all in a manner that relies on statements about what lies behind the data, e.g. whether nodes were more or less likely to be connected to begin with. A majority of conclusions reached would be severely undermined if one would discover that the underlying network is in fact fully random. This means that these analyses are at a grave peril when using purely descriptive methods, since they are likely to be overfitting the data — i.e. confusing randomness with underlying structure.
References
Webmentions3
(Nothing yet)
3 Webmention is a standardized decentralized mechanism for conversations and interactions across the web.
Comments
(Comments may be moderated.)
If you reply to this post, it will show as a comment below the original article.