Inferring, explaining, and compressing
Inferential approaches to community detection (see [2] for a detailed introduction) are designed to provide explanations for network data in a principled manner. They are based on the formulation of generative models that include the notion of community structure in the rules of how the edges are placed. More formally, they are based on the definition of a likelihood \(P({\boldsymbol A}|{\boldsymbol b})\) for the network \({\boldsymbol A}\) conditioned on a partition \({\boldsymbol b}\), and the inference is obtained via the posterior distribution, according to Bayes' rule, i.e.
\[P({\boldsymbol b}|{\boldsymbol A}) = \frac{P({\boldsymbol A}|{\boldsymbol b})P({\boldsymbol b})}{P({\boldsymbol A})}, \tag{1}\]
where \(P({\boldsymbol b})\) is the prior probability for a partition \({\boldsymbol b}\). Overwhelmingly, the models used for this purpose are variations of the stochastic block model (SBM) [3], where in addition to the node partition, it takes the probability of edges being placed between the different groups as an additional set of parameters. A particularly expressive variation is the degree-corrected SBM (DC-SBM) [4], with a marginal likelihood given by [5].
\[P({\boldsymbol A}|{\boldsymbol b}) = \sum_{\boldsymbol e, \boldsymbol k}P({\boldsymbol A}|{\boldsymbol k},{\boldsymbol e},{\boldsymbol b})P({\boldsymbol k}|{\boldsymbol e},{\boldsymbol b})P({\boldsymbol e}|{\boldsymbol b}), \tag{2}\]
where \({\boldsymbol e}=\{e_{rs}\}\) is a matrix with elements \(e_{rs}\) specifying how many edges go between groups \(r\) and \(s\), and \(\boldsymbol k=\{k_i\}\) are the degrees of the nodes. Therefore, this model specifies that, conditioned on a partition \({\boldsymbol b}\), first the edge counts \({\boldsymbol e}\) are sampled from a prior distribution \(P({\boldsymbol e}|{\boldsymbol b})\), followed by the degrees from the prior \(P({\boldsymbol k}|{\boldsymbol e},{\boldsymbol b})\), and finally the network is wired together according to the probability \(P({\boldsymbol A}|{\boldsymbol k},{\boldsymbol e},{\boldsymbol b})\), which respects the constraints given by \(\boldsymbol k\), \({\boldsymbol e}\), and \({\boldsymbol b}\). See Figure 1 (a) for a illustration of this process.
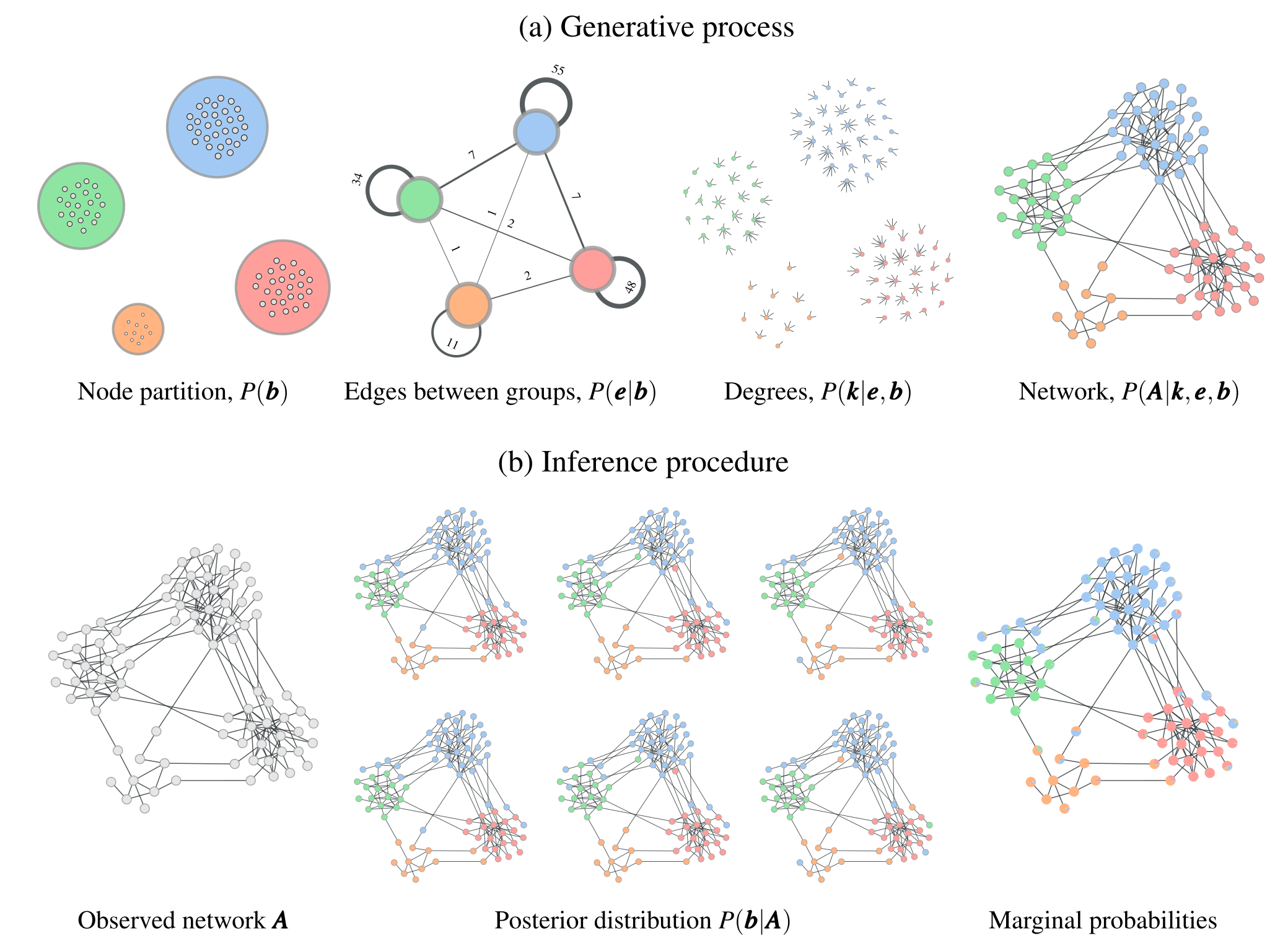
This model formulation includes fully random networks as the special case when we have a single group. Together with the Bayesian approach, the use of this model will inherently favor a more parsimonious account of the data, whenever it does not warrant a more complex description — amounting to a formal implementation of Occam's razor. This is best seen by making a formal connection with information theory, and noticing that we can write the numerator of Equation 1 as
\[P({\boldsymbol A}|{\boldsymbol b})P({\boldsymbol b}) = 2^{-\Sigma({\boldsymbol A},{\boldsymbol b})},\]
where the quantity \(\Sigma({\boldsymbol A},{\boldsymbol b})\) is known as the description length [6] of the network. It is computed as:1
1 Note that the sum in Equation 2 vanishes because only one term is non-zero given a fixed network \({\boldsymbol A}\).
\[\Sigma({\boldsymbol A},{\boldsymbol b}) = \underset{\mathcal{D}({\boldsymbol A}|{\boldsymbol k},{\boldsymbol e},{\boldsymbol b})}{\underbrace{-\log_2P({\boldsymbol A}|{\boldsymbol k},{\boldsymbol e},{\boldsymbol b})}}\, \underset{\mathcal{M}({\boldsymbol k},{\boldsymbol e},{\boldsymbol b})}{\underbrace{-\log_2P({\boldsymbol k}|{\boldsymbol e},{\boldsymbol b}) - \log_2 P({\boldsymbol e}|{\boldsymbol b}) - \log_2P({\boldsymbol b})}}. \tag{3}\]
The second set of terms \(\mathcal{M}({\boldsymbol k},{\boldsymbol e},{\boldsymbol b})\) in the above equation quantifies the amount of information in bits necessary to encode the parameters of the model 2. The first term \(\mathcal{D}({\boldsymbol A}|{\boldsymbol k},{\boldsymbol e},{\boldsymbol b})\) determines how many bits are necessary to encode the network itself, once the model parameters are known. This means that if Bob wants to communicate to Alice the structure of a network \({\boldsymbol A}\), he first needs to transmit \(\mathcal{M}({\boldsymbol k},{\boldsymbol e},{\boldsymbol b})\) bits of information to describe the parameters \({\boldsymbol b}\), \({\boldsymbol e}\), and \({\boldsymbol k}\), and then finally transmit the remaining \(\mathcal{D}({\boldsymbol A}|{\boldsymbol k},{\boldsymbol e},{\boldsymbol b})\) bits to describe the network itself. Then, Alice will be able to understand the message by first decoding the parameters \(({\boldsymbol k},{\boldsymbol e},{\boldsymbol b})\) from the first part of the message, and using that knowledge to obtain the network \({\boldsymbol A}\) from the second part, without any errors.
2 If a value \(x\) occurs with probability \(P(x)\), this means that in order to transmit it in a communication channel we need to answer at least \(-\log_2P(x)\) yes-or-no questions to decode its value exactly. Therefore we need to answer one yes-or-no question for a value with \(P(x)=1/2\), zero questions for \(P(x)=1\), and \(\log_2N\) questions for uniformly distributed values with \(P(x)=1/N\). This value is called “information content”, and essentially measures the degree of “surprise” when encountering a value sampled from a distribution. See [7] for a thorough but accessible introduction to information theory and its relation to inference.
What the above connection shows is that there is a formal equivalence between inferring the communities of a network and compressing it. This happens because finding the most likely partition \({\boldsymbol b}\) from the posterior \(P({\boldsymbol b}|{\boldsymbol A})\) is equivalent to minimizing the description length \(\Sigma({\boldsymbol A},{\boldsymbol b})\) used by Bob to transmit a message to Alice containing the whole network.
Data compression amounts to formal implementation of Occam's razor because it penalizes models that are too complicated: if we want to describe a network using many communities, then the model part of the description length \(\mathcal{M}({\boldsymbol k},{\boldsymbol e},{\boldsymbol b})\) will be large, and Bob will need many bits to transmit the model parameters to Alice. However, increasing the complexity of the model will also reduce the first term \(\mathcal{D}({\boldsymbol A}|{\boldsymbol k},{\boldsymbol e},{\boldsymbol b})\), since there are fewer networks that are compatible with the bigger set of constraints, and hence Bob will need a shorter second part of the message to convey the network itself once the parameters are known. Compression (and hence inference), therefore, is a balancing act between model complexity and quality of fit, where an increase in the former is only justified when it results in an even larger increase of the second, such that the total description length is minimized.
The reason why the compression approach avoids overfitting the data is due to a powerful fact from information theory, known as Shannon's source coding theorem [8], which states that it is impossible to compress data sampled from a distribution \(P(x)\) using fewer bits per symbol than the entropy of the distribution, \(H=-\sum_xP(x)\log_2P(x)\). In our context, this means that it is impossible, for example, to compress a fully random network using a SBM with more than one group.3 This means, for example, that when encountered with an example like in the figure we considered in the previous blog post, inferential methods will detect a single community comprising all nodes in the network, since any further division does not provide any increased compression, or equivalently, no augmented explanatory power. From the inferential point of view, a partition like in the previous figure (b) overfits the data, since it incorporates irrelevant random features — a.k.a. “noise” — into its description.
3 More accurately, this becomes impossible only when the network becomes asymptotically infinite; for finite networks the probability of compression is only vanishingly small.
In Figure 2 (a) is shown an example of the results obtained with an inferential community detection algorithm, for a network sampled from the SBM. As shown in Figure 2 (b), the obtained partitions are still valid when carried over to an independent sample of the model, because the algorithm is capable of separating the general underlying pattern from the random fluctuations. As a consequence of this separability, this kind of algorithm does not find communities in fully random networks, which are composed only of “noise.”

Role of inferential approaches in community detection
Inferential approaches based on the SBM have an old history, and were introduced for the study of social networks in the early 80's [3]. But despite such an old age, and having appeared repeatedly in the literature over the years (also under different names in other contexts), they entered the mainstream community detection literature rather late, arguably after the influential paper by Karrer and Newman that introduced the DC-SBM [4] in 2011, at a point where descriptive approaches were already dominating. However, despite the dominance of descriptive methods, the existence of inferential criteria was already long noticeable. In fact, in a well-known attempt to systematically compare the quality of a variety of descriptive community detection methods, the authors of [9] proposed the now so-called LFR benchmark, offered as a more realistic alternative to the simpler Newman-Girvan benchmark [10] introduced earlier. Both are in fact generative models, essentially particular cases of the DC-SBM, containing a “ground truth” community label assignment, against which the results of various algorithms are supposed to be compared. Clearly, this is an inferential evaluation criterion, although, historically, virtually all of the methods compared against that benchmark are descriptive in nature [11] (these studies were conducted mostly before inferential approaches had gained more traction). The use of such a criterion already betrays that the answer to the litmus test considered in the previous post would be “yes,” and therefore descriptive approaches are fundamentally unsuitable for the task. In contrast, methods based on statistical inference are not only more principled, but in fact provably optimal in the inferential scenario, in the sense that all conceivable algorithms can obtain either equal or worse performance, but none can do better [12].
The conflation one often finds between descriptive and inferential goals in the literature of community detection likely stems from the fact that while it is easy to define benchmarks in the inferential setting, it is substantially more difficult to do so in a descriptive setting. Given any descriptive method (modularity maximization, Infomap, Markov stability, etc.) it is usually problematic to determine for which network these methods are optimal (or even if one exists), and what would be a canonical output that would be unambiguously correct. In fact, the difficulty with establishing these fundamental references already serve as evidence that the task itself is ill-defined. On the other hand, taking an inferential route forces one to start with the right answer, via a well-specified generative model that articulates what the communities actually mean with respect to the network structure. Based on this precise definition, one then derives the optimal detection method by employing Bayes' rule.
It is also useful to observe that inferential analyses of aspects of the network other than directly its structure might still be only descriptive of the structure itself. A good example of this is the modelling of dynamics that take place on a network, such as a random walk. This is precisely the case of the Infomap method, which models a simulated teleporting random walk on a network in an inferential manner, using for that a division of the network into groups. While this approach can be considered inferential with respect to an artificial dynamics, it is still only descriptive when it comes to the actual network structure (and will suffer the same problems, such a finding communities in fully random networks). Communities found in this way could be useful for particular tasks, such as to identify groups of nodes that would be similarly affected by a diffusion process. This could be used, for example, to prevent or facilitate the diffusion by removing or adding edges between the identified groups. In this setting, the answer to the litmus test would also be “no”, since what is important is how the network “is” (i.e. how a random walk behaves on it), not how it came to be, or if its features are there by chance alone. Once more, the important issue to remember is that the groups identified in this manner cannot be interpreted as having any explanatory power about the network structure itself, and cannot be used reliably to extract inferential conclusions from it. We are firmly in a descriptive, not inferential setting with respect to the network structure.
Another important difference between inferential and descriptive approaches is worth mentioning. Descriptive approaches are tied to very particular contexts, and cannot be directly compared to one another. This has caused great consternation in the literature, since there is a vast number of such methods, and little robust methodology on how to compare them. Indeed, why should we expect that the modules found by optimizing task scheduling should be comparable to those that optimize the description of a dynamics? In contrast, inferential approaches all share the same underlying context: they attempt to explain the network structure; they vary only in how this is done. They are, therefore, amenable to principled model selection procedures, designed to evaluate which is the most appropriate fit for any particular network, even if the models used operate with very different parametrizations. In this situation, the multiplicity of different models available becomes a boon rather than a hindrance, since they all contribute to a bigger toolbox we have at our disposal when trying to understand empirical observations.
Finally, inferential approaches offer additional advantages that make them more suitable as part a scientific pipeline. In particular, they can be naturally extended to accommodate measurement uncertainties [13] — an unavoidable property of empirical data, which descriptive methods almost universally fail to consider. This information can be used not only to propagate the uncertainties to the community assignments [14] but also to reconstruct the missing or noisy measurements of the network itself [15]. Furthermore, inferential approaches can be coupled with even more indirect observations such as time-series on the nodes [16], instead of a direct measurement of the edges of the network, such that the network itself is reconstructed, not only the community structure [17]. All these extensions are possible because inferential approaches give us more than just a division of the network into groups; they give us a model estimate of the network, containing insights about its formation mechanism.
Behind every description there is an implicit generative model
From a purely mathematical perspective, there is actually no formal distinction between descriptive and inferential methods, because every descriptive method can be mapped to an inferential one, according to some implicit model. Therefore, whenever we are attempting to interpret the results of a descriptive community detection method in an inferential way — i.e. make a statement about how the network came to be — we cannot in fact avoid making implicit assumptions about the model generating process that lies behind it. (At first this statement seems to undermine the distinction we have been making between descriptive and inferential methods, but in fact this is not the case, as we will see below.)
It is not difficult to demonstrate that it is possible to formulate any conceivable community detection method as a particular inferential method. Let us consider an arbitrary quality function
\[W({\boldsymbol A}, {\boldsymbol b}) \in \mathbb{R}\]
which is used to perform community detection via the optimization
\[{\boldsymbol b}^* = \underset{\boldsymbol b}{\operatorname{argmax}}\; W({\boldsymbol A}, {\boldsymbol b}). \tag{4}\]
We can then interpret the quality function \(W({\boldsymbol A}, {\boldsymbol b})\) as the “Hamiltonian” of a posterior distribution
\[P({\boldsymbol b}|{\boldsymbol A}) = \frac{\mathrm{e}^{\beta W({\boldsymbol A},{\boldsymbol b})}}{Z({\boldsymbol A})},\]
with normalization \(Z({\boldsymbol A})=\sum_{\boldsymbol b}\mathrm{e}^{\beta W({\boldsymbol A},{\boldsymbol b})}\). By making \(\beta\to\infty\) we recover the optimization of Equation 4, or we may simply try to find the most likely partition according to the posterior, in which case \(\beta>0\) remains an arbitrary parameter. Therefore, employing Bayes' rule in the opposite direction, we obtain the following effective generative model:
\[\begin{aligned} \begin{aligned} P({\boldsymbol A}|{\boldsymbol b}) &= \frac{P({\boldsymbol b}|{\boldsymbol A})P({\boldsymbol A})}{P({\boldsymbol b})},\\ &= \frac{\mathrm{e}^{\beta W({\boldsymbol A},{\boldsymbol b})}}{Z({\boldsymbol A})}\frac{P({\boldsymbol A})}{P({\boldsymbol b})}, \end{aligned} \end{aligned}\]
where \(P({\boldsymbol A}) = \sum_{\boldsymbol b}P({\boldsymbol A}|{\boldsymbol b})P({\boldsymbol b})\) is the marginal distribution over networks, and \(P({\boldsymbol b})\) is the prior distribution for the partition. Due to the normalization of \(P({\boldsymbol A}|{\boldsymbol b})\) we have the following constraint that needs to be fulfilled:
\[\sum_{\boldsymbol A}\frac{\mathrm{e}^{\beta W({\boldsymbol A},{\boldsymbol b})}}{Z({\boldsymbol A})}P({\boldsymbol A}) = P({\boldsymbol b}). \tag{5}\]
Therefore, not all choices of \(P({\boldsymbol A})\) and \(P({\boldsymbol b})\) are compatible with the posterior distribution and the exact possibilities will depend on the actual shape of \(W({\boldsymbol A},{\boldsymbol b})\). However, one choice that is always possible is
\[P({\boldsymbol A}) = \frac{Z({\boldsymbol A})}{\Xi},\qquad P({\boldsymbol b}) = \frac{\Omega({\boldsymbol b})}{\Xi},\]
with \(\Omega({\boldsymbol b})=\sum_{\boldsymbol A}\mathrm{e}^{\beta W({\boldsymbol A},{\boldsymbol b})}\) and \(\Xi=\sum_{\boldsymbol A,\boldsymbol b}\mathrm{e}^{\beta W({\boldsymbol A},{\boldsymbol b})}\). Taking this choice leads to the effective generative model
\[P({\boldsymbol A}|{\boldsymbol b}) = \frac{\mathrm{e}^{\beta W({\boldsymbol A},{\boldsymbol b})}}{\Omega({\boldsymbol b})}.\]
Therefore, inferentially interpreting a community detection algorithm with a quality function \(W({\boldsymbol A},{\boldsymbol b})\) is equivalent to assuming the generative model \(P({\boldsymbol A}|{\boldsymbol b})\) and prior \(P({\boldsymbol b})\) above. Furthermore, this also means that any arbitrary community detection algorithm implies a description length4 given (in nats) by
4 The description length of Equation 6 is only valid if there are no further parameters in the quality function \(W({\boldsymbol A},{\boldsymbol b})\) other than \({\boldsymbol b}\) that are being optimized.
\[\Sigma({\boldsymbol A},{\boldsymbol b}) = -\beta W({\boldsymbol A},{\boldsymbol b}) + \ln\sum_{\boldsymbol A',{\boldsymbol b}'}\mathrm{e}^{\beta W({\boldsymbol A}',{\boldsymbol b}')}. \tag{6}\]
What the above shows is that there is no such thing as a “model-free” community detection method, since they are all equivalent to the inference of some generative model. The only difference to a direct inferential method is that in that case the modelling assumptions are made explicitly, inviting rather than preventing scrutiny. Most often, the effective model and prior that are equivalent to an ad hoc community detection method will be difficult to interpret, justify, or even compute.
Furthermore there is no guarantee that the obtained description length of Equation 6 will yield a competitive or even meaningful compression. In particular, there is no guarantee that this effective inference will not overfit the data. Although we mentioned in the previous section that inference and compression are equivalent, the compression achieved when considering a particular generative model is constrained by the assumptions encoded in its likelihood and prior. If these are poorly chosen, no actual compression might be achieved, for example when comparing to the one obtained with a fully random model. This is precisely what happens with descriptive community detection methods: they overfit because their implicit modelling assumptions do not accommodate the possibility that a network may be fully random, or contain a balanced mixture of structure and randomness.
Since we can always interpret any community detection method as inferential, is it still meaningful to categorize some methods as descriptive? Arguably yes, because directly inferential approaches make their generative models and priors explicit, while for a descriptive method we need to extract them from back-engineering. Explicit modelling allows us to make judicious choices about the model and prior that reflect the kinds of structures we want to detect, relevant scales or lack thereof, and many other aspects that improve their performance in practice, and our understanding of the results. With implicit assumptions we are “flying blind”, relying substantially on serendipity and trial-and-error — not always with great success.
It is not uncommon to find criticisms of inferential methods due to a perceived implausibility of the generative models used — such as the conditional independence of the placement of the edges present in the SBM — although these assumptions are also present, but only implicitly, in other methods, like modularity maximization (see [1]).
The above inferential interpretation is not specific to community detection, but is in fact valid for any learning problem. The set of explicit or implicit assumptions that must come with any learning algorithm is called an “inductive bias”. An algorithm is expected to function optimally only if its inductive bias agrees with the actual instances of the problems encountered. It is important to emphasize that no algorithm can be free of an inductive bias, we can only chose which intrinsic assumptions we make about how likely we are to encounter a particular kind of data, not whether we are making an assumption. Therefore, it is particularly problematic when a method does not articulate explicitly what these assumptions are, since even if they are hidden from view, they exist nonetheless, and still need to be scrutinized and justified. This means we should be particularly skeptical of the impossible claim that a learning method is “model free”, since this denomination is more likely to signal an unwillingness to expose the underlying modelling assumptions, which could potentially be revealed as unappealing and fragile when eventually forced to come under scrutiny.
References
Webmentions5
(Nothing yet)
5 Webmention is a standardized decentralized mechanism for conversations and interactions across the web.
Comments